Function describing the standard deviation of the measurement error in dependence of the measured value \(y\):
Arguments
- y
The magnitude of the observed value
- sigma_low
The asymptotic minimum of the standard deviation for low observed values
- rsd_high
The coefficient describing the increase of the standard deviation with the magnitude of the observed value
Details
$$\sigma = \sqrt{ \sigma_{low}^2 + y^2 * {rsd}_{high}^2}$$ sigma = sqrt(sigma_low^2 + y^2 * rsd_high^2)
This is the error model used for example by Werner et al. (1978). The model proposed by Rocke and Lorenzato (1995) can be written in this form as well, but assumes approximate lognormal distribution of errors for high values of y.
References
Werner, Mario, Brooks, Samuel H., and Knott, Lancaster B. (1978) Additive, Multiplicative, and Mixed Analytical Errors. Clinical Chemistry 24(11), 1895-1898.
Rocke, David M. and Lorenzato, Stefan (1995) A two-component model for measurement error in analytical chemistry. Technometrics 37(2), 176-184.
Ranke J and Meinecke S (2019) Error Models for the Kinetic Evaluation of Chemical Degradation Data. Environments 6(12) 124 doi:10.3390/environments6120124 .
Examples
times <- c(0, 1, 3, 7, 14, 28, 60, 90, 120)
d_pred <- data.frame(time = times, parent = 100 * exp(- 0.03 * times))
set.seed(123456)
d_syn <- add_err(d_pred, function(y) sigma_twocomp(y, 1, 0.07),
reps = 2, n = 1)[[1]]
f_nls <- nls(value ~ SSasymp(time, 0, parent_0, lrc), data = d_syn,
start = list(parent_0 = 100, lrc = -3))
library(nlme)
f_gnls <- gnls(value ~ SSasymp(time, 0, parent_0, lrc),
data = d_syn, na.action = na.omit,
start = list(parent_0 = 100, lrc = -3))
if (length(findFunction("varConstProp")) > 0) {
f_gnls_tc <- update(f_gnls, weights = varConstProp())
f_gnls_tc_sf <- update(f_gnls_tc, control = list(sigma = 1))
}
f_mkin <- mkinfit("SFO", d_syn, error_model = "const", quiet = TRUE)
f_mkin_tc <- mkinfit("SFO", d_syn, error_model = "tc", quiet = TRUE)
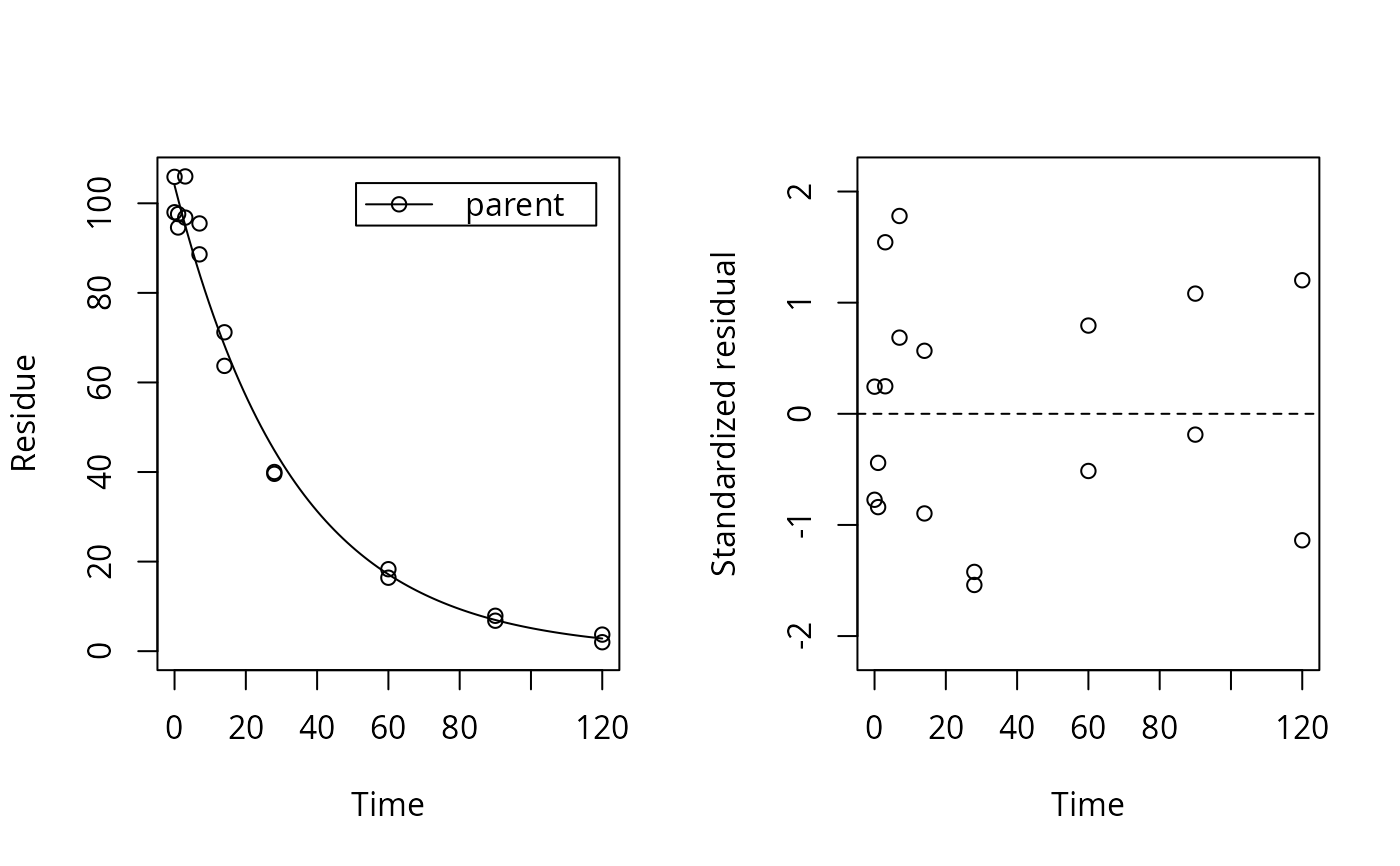
plot_res(f_mkin_tc, standardized = TRUE)
 AIC(f_nls, f_gnls, f_gnls_tc, f_gnls_tc_sf, f_mkin, f_mkin_tc)
#> df AIC
#> f_nls 3 114.4817
#> f_gnls 3 114.4817
#> f_gnls_tc 5 103.6447
#> f_gnls_tc_sf 4 101.6447
#> f_mkin 3 114.4817
#> f_mkin_tc 4 101.6446
AIC(f_nls, f_gnls, f_gnls_tc, f_gnls_tc_sf, f_mkin, f_mkin_tc)
#> df AIC
#> f_nls 3 114.4817
#> f_gnls 3 114.4817
#> f_gnls_tc 5 103.6447
#> f_gnls_tc_sf 4 101.6447
#> f_mkin 3 114.4817
#> f_mkin_tc 4 101.6446